Short Communication 
 Creative Commons, CC-BY
Creative Commons, CC-BY
Porcine Respiratory and Reproductive Syndrome Virus (PRRSV): Infection of Piglets
*Corresponding author: Július Rajčáni, RT-Europe Non-profit Research Centre, Vár tér 2/E, Mosonmagyaróvár, H-9200, Hungary
Received: November 13, 2021; Published: November 29, 2021
DOI: 10.34297/AJBSR.2021.14.002049
Short Communication
The porcine respiratory and reproduction syndrome virus (PRRSV) forms small, enveloped particles (50-65 nm in diameter), which contain a relatively large (approximately 15 kb long) single strand RNA genome. More precisely, the viral RNA (vRNA) is a positive-sense, single-stranded molecule with a terminal cap at its 5´-end and a poly-A repeat at the 3´sd-end. The vRNA is copied as whole by means of a full-length complementary (negative sense) intermediate strand. The resulting minus sense subgenomic (sg) RNA is then used as template for the synthesis of the additional vRNAs. The enzyme involved is the RNA-dependent RNA polymerase (RdRp), also termed non-structural protein 9 (nsp9), encoded by the virus. The vRNA contains 2 long open reading frames (namely ORF1a and ORF1b); all the 14 non-structural proteins (nsps) are formed by cleavage of 2 large polyproteins, the coding sequence of which streches nearly over 75% of the total genome length [1]. The ORF1b sequence encodes 7 structural proteins (M, N and E) out of which 4 are glycoproteins, namely GP2a, GP3, GP4, GP5. Based on the structure of vRNA decribed above, PRRSV has been classified into family Arteriviridae, along with equine arteriitis virus and/ or the lactate dehydrogenase elevating virus of mice. Based on the described structure of vRNA, the PRRSV has been classified into family Arteriviridae, along with equine arteriitis virus and/or lactate dehydrogenase elevating virus of mice.
Each coding sequence of the vRNA has a short transcription regulating sequence (TRS), which is strictly conserved and strictly located preceeding its position. In the coarse of transcription, the replicase complex runs across the TRS to copy the specific gene sequence. This way a complementary (minus) strand is being formed, in which the TRS body is positioned at the 5´-UTR end. By forming a base pair with the leader sequence, the TRS sequence helps the polymerase complex to restart the RNA synthesis. The resulting minus sense subgenomic (sg) RNA is used as template for the synthesis of the sg RNAs [2].
The two prototype strains which were isolated either in US (VR2332) or in Europe (Lelystadt) when compared to each other have revealed serologic as well as genomic differences [3]. Experimental infection with PRRSV has been successful in both, the newborn as well as in 3-week-old piglets, even with different isolates. The key event causing development of disease is the involvement of alveolar macrophages, which represent an important target for virus replication as well as its spread [4]. To date, two macrophagespecific molecules are known as PRRSV entry mediators: the siglet sialoadhesin and a scavenger receptor (CD163) [5]. The PRRSV induced pneumonia is characterized by thickening of interalveolar septa due to infiltration with macrophages and occasional the presence of cell debris within the alveolar cavity [6]. The infected lung tissue shows a typical picture of usual interstitial pneumonia (UIP) revealing a wide thickening of the septi in between alveoli, which may be infiltrated by mononuclear cells, mainly lymphocytes. Occasionally the capillaries of the septi are widened as well, along with focal bleeding from injured endothelium cells. High power view of such area shows that the interstitial infiltrate consists mainly of lymphocytes [7]. Proliferating alveolar pneumocytes of type II can be also found along with hyperplastic peribronchial lymphatic tissue [8]. The severity of lung lesions vary from mild to extensive.
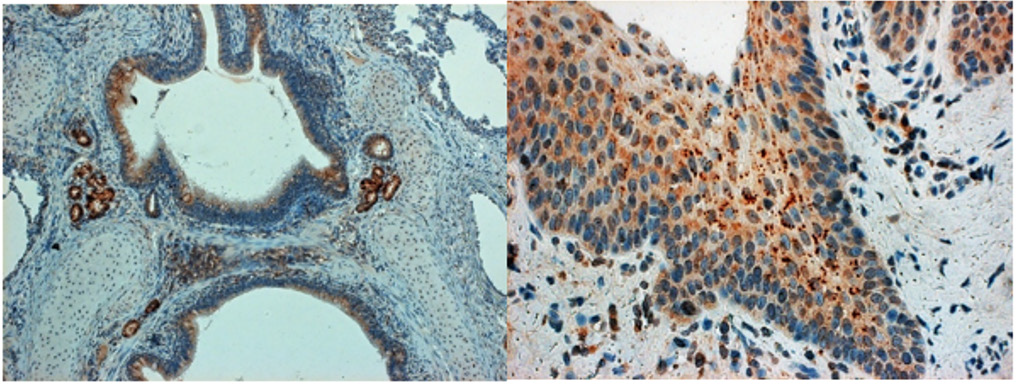
The different genotypes (namely the Type 2 North American PRRSV induces more severe respiratory disease than type 1 European virus). As described, PRRSV replicates predominantly in the lung alveolar macrophages, can induce prolonged viremia, and cause persistent infections lasting for months after initial infection. PRRSV strongly modulates the host’s immune responses and changes the host’s gene expression. The N-protein of PRRSV could be seen in the bronchial ciliary epithelium cells of piglets who developed UIP along with less frequently positive squamous epithelium at pharyngeal and/or tonsillar areas (Figure 1).
References
- Conzelmann KK, N Visser, P Van Woensel, HJ Thiel (1993) Molecular characterization of porcine reproductive and respiratory syndrome virus, a member of the arterivirus group. Virology 193(1): 329-339.
- Pasternak A O, E van den Born, W J Spaan, E J Snijder (2001) Sequence requirements for RNA strand transfer during nidovirus discontinuous subgenomic RNA synthesis. Embo J 20(24): 7220-7228.
- Katz JB, Shafer AL, Eernisse, KA, Landgraf, JG, Nelson EA (1995) Antigenic differences between European and American isolates of procine reproductive and respiratory syndrome virus (PRRSV) are encoded by the carboxy-terminal portion of viral open reading frame 3. Vet Microbiol 44(1): 65-76.
- Lawson S R, K D Rossow, J E Collins, D A Benfield, R R Rowland (1997) Porcine reproductive and respiratory syndrome virus infection of gnotobiotic pigs: sites of virus replication and co-localization with MAC-387 staining at 21 days post-infection. Virus Res 51(2):105-113.
- Van Gorp H, W Van Breedam, P L Delputte, HJ Nauwynck (2008) Sialoadhesin and CD163 join forces during entry of the porcine reproductive and respiratory syndrome virus. J Gen Virol 89(Pt 12): 2943-2953.
- E Rubin, JL Farber (Eds.), Pathology 2nd Edn (1994) The respiratory system. Lippincott Company, Philadelphia. pp. 556-617.
- Rossow KD (1998) Porcine Reproductive Respiratory Syndrome. Review Article. Vet Pathol 35: 1-20.
- Rossow KD, Benfield DA, Goyal SM, Nerlson EA, Christopher Hennings J, et al. (1996) Chronical immunohistochemical detection of the porcine reproductive and respiratory syndrome virus infection in gnotobiotic pigs. Vet Pathol 33(5): 551-556.



 We use cookies to ensure you get the best experience on our website.
We use cookies to ensure you get the best experience on our website.