Research Article 
 Creative Commons, CC-BY
Creative Commons, CC-BY
Computational Pharmacokinetics of Nanopharmacophores: Integrating Evolutionary Algorithms in In Silico Simulations
*Corresponding author:Valentin V Fursov, N.I. Pirogov Russian National Research Medical University, Russia
Received:February 02, 2026; Published:February 11, 2026
DOI: 10.34297/AJBSR.2026.30.003885
Abstract
Background: With the increasing complexity of optimization problems in the scientific environment, evolutionary algorithms are becoming more and more popular. One area in which such algorithms are gaining popularity is pharmacokinetics.
Methods: In this study, a new evolutionary algorithm of interacting countries inspired by the classical genetic algorithm, island algorithm and migration algorithm is proposed. The algorithm is applied to find the unknown parameters of a five-compartment pharmacokinetic model describing the pharmacokinetics of porphyrin-fullerene nanoparticles (PMC16) in the body when administered once intravenously.
Results: The algorithm enables significant improvement in the accuracy of in silico modeling results compared to the pharmacokinetic curves obtained from PMC16 distribution in heart, liver, and brain tissues in vivo.
Conclusion: The results obtained with using the algorithm prove its efficiency and applicability in this field of in silico modeling of pharmacokinetics of porphyrin-fullerene nano pharmacophores.
Keywords:Models of population dynamics, Complex systems models, Biosystem theoretical models, In silico, Evolutionary models, Genetic algorithm, Porphyrin-fullerene nanocationites, Neuroprotectors
Introduction
The complexity of problems solved by the scientific community has increased over time in scientific fields such as robotics, bioinformatics, decision-making, machine learning, and others. For most of these problems, the optimal solution is non-obvious and difficult to achieve. To solve them, an approach based on Darwinian natural evolution has been proposed, which has been given the name Evolutionary Computing (EC) [1-2].
ECs involve a series of algorithms called Evolutionary Algorithms (EAs). EAs model the evolution of a population to find optimal solutions to complex problems. The main application of EAs is related to problems for which the use of a simple deterministic algorithm is impossible or may lead to inadequate results. For example, it is known that simple algorithms cannot find the global extremum when the objective function of optimisation is complex. In recent years, the subject of EAs has become increasingly popular, especially in the context of their application to solve complex practical optimisation problems in different areas of economy, industry, and medicine [3-7].
Currently, evolutionary algorithms are increasingly applied in the field of pharmacokinetics, including the search for covariance models in population pharmacokinetic analysis [8,9]. Separately, these algorithms are widely used to search for unknown parameters of pharmacokinetic models [10,11]. Thus, in multi-compartment models, where different organs are represented as chambers connected to each other [12], parameters affecting the rate of drug transport between organs can be estimated.
The recent discovery of the biophysical control of the energy production process in mitochondria through the so-called magnesium-25 magnetic isotope effect prompted the development of a pharmacologically suitable [Mg2+] ion nanocarrier. It is the magnetic isotope of magnesium [25Mg2+] (nuclear spin +5/2, magnetic moment 0.85 Bohr magneton, fraction in the natural isotope mixture 11 %), which has been shown to be a specific hyper activator of most [Mg2+]-dependent reactions of ATP synthesis in the cell. In contrast to the non-magnetic isotopes [24Mg2+] and [26Mg2+], it is the only magnetic isotope of magnesium. It has been stated that hyperactivation of energy metabolism by [25Mg2+] ions requires negligible amounts of these ions and occurs even in conditions of deep tissue hypoxia [13].
This biophysical phenomenon could be used to correct the reduction of ATP formation in tissues under hypoxia of any nature. Earlier, a new pharmaceutical preparation based on the porphyrincontaining fullerene “ball” C60 has been proposed [14,15]. This novel drug is a monoadduct of an iron-containing porphyrin with the classical buckminsterfullerene, buckminsterfullerene(C60)- 2-(butadiene-1-yl)-tetra-(o-γ-aminobutyryl-o-fthalyl)- ferroporphyrin. It was named porphyllerene-MC16, or PMC16. The particles are low-toxic and exhibit notable cationic properties.
Some drugs can suppress oxygenating ADP phosphorylation in mitochondria and cause hypoxia. Nano quantities of [25 Mg2+] could accelerate phosphorylation pathways up to 2.5 times [15]. Mg isotopes substitute each other in the active site of creatine kinase owing to endoosmotic pressure [13]. This promotes the magnetic isotope effect of [25 Mg2+], which is an important hyperactivating element of the magnesium-dependent control of ATP synthesis [15-18]. One can conclude, that the targeted delivery of the aforementioned magnetic isotope to hypoxia-affected cells or tissues is a really important pharmacological problem. Nanoparticles capable of [Mg2+] ion exchange may be a suitable vehicle for such delivery [13].
It should be noted that the creation of in silico models of pharmacokinetics of targeting and selective accumulation of PMC16 line nanoparticles is extremely important for optimising further research in this field and has all the necessary novelty and relevance. Speaking of the direct and explicit practical benefits that are expected to be derived from the appropriate use of mathematical modelling in determining the preclinical study design of antihypoxic drugs, it should be noted that such modelling is an essential requirement for optimising the study design of all new drugs.
This paper will discuss the application of a new evolutionary algorithm, the interacting countries algorithm [19], to solve the problem of finding the parameters of a pharmacokinetic model of intravenous administration of porphyrin-fullerene PMC16 nanoparticles releasing [25 Mg2+]. The aim of the work was to improve the agreement of the model with experimental data of PMC16 distribution in organs in comparison with the previous work [20]. This article will consider the application of a new evolutionary algorithm – the algorithm of interacting countries [19] to solve the problem of determining the parameters of a pharmacokinetic model of intravenous administration of porphyrin-fullerene PMC16 nanoparticles releasing [25 Mg2+]. The aim of the work was to improve the agreement of the model with experimental data of PMC16 distribution in organs in comparison with the previous work [20].
Materials and Methods
The Algorithm of Interacting Countries
Before presenting the point of the algorithm, it should be noted that in this context, “country” means a group of N individuals (solutions). The proposed algorithm includes a sequence of actions performed either by the whole country or by separate individuals within it. All actions performed within the framework of the algorithm can be conditionally divided into natural and specially created ones. Let us start the description of the algorithm with the description of natural actions; there are only two of them – reproduction and extinction.
Reproduction
By analogy with the genetic algorithm, in the algorithm of interacting countries, all individuals within the country are trying to leave their legacy by reproduction. At the same time, it is worth noting that the algorithms for the formation of a child based on two (or more) parents do not play a role in this case and can be selected during implementation, as well as the algorithm for selecting parental individuals from the population. However, the algorithm determines the number of pairs producing offspring for each individual country. This number for the i-th country for the minimisation process is determined by the formula (1):
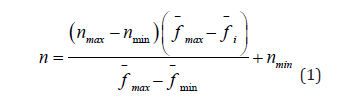
where n is the required number of pairs, nmax is the maximum
number of pairs, nmin is the minimum number of pairs,
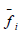 is the
average value of the objective function for the country,
is the
average value of the objective function for the country,
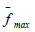 is
the maximum average value of the objective function among
all countries,
is
the maximum average value of the objective function among
all countries, 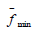 is the minimum average value of the objective
function among all countries. The found n is rounded to the nearest
integer. This mechanism ensures an increase in the number of
offspring in more adapted countries and a decrease in less adapted
ones. It can be considered logical that in societies with a higher level
of well-being and greater resources, more offspring will be born,
similar to animals in nature, where more prosperous conditions
ensure a higher birth rate. When a new individual is formed, each of
its genes (an independent variable) undergoes mutation. Thus, the
i-th gene can be calculated using the formulas (2-3):
is the minimum average value of the objective
function among all countries. The found n is rounded to the nearest
integer. This mechanism ensures an increase in the number of
offspring in more adapted countries and a decrease in less adapted
ones. It can be considered logical that in societies with a higher level
of well-being and greater resources, more offspring will be born,
similar to animals in nature, where more prosperous conditions
ensure a higher birth rate. When a new individual is formed, each of
its genes (an independent variable) undergoes mutation. Thus, the
i-th gene can be calculated using the formulas (2-3):
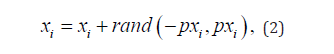
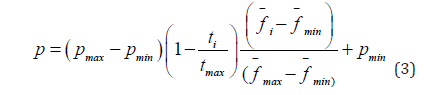
where p is the mutational deviation, pmax is the maximum mutational deviation, pmin is the minimum mutational deviation, ti is the current iteration of the algorithm, tmax is the maximum possible iteration of the algorithm. The mutation mechanism allows you to diversify the set of individuals in a country to reduce the chance of degeneration. In addition, it can be noted that the deviation from the parent individuals in the first iterations will be much larger than in the last, which will allow you to quickly find the area to find the optimum, this idea is inspired by the weed algorithm. Also note that for better adapted countries, offspring will mutate less than for less adapted ones, which will allow the more adapted to localize in the area they have found and the less adapted, on the contrary, to move away from the area where they are.
Extinction
The second natural action, which happens to countries, is extinction. Each iteration of the algorithm removes m worst individuals from each country based on the value of their objective function. The value of m can be found by the formula (4):
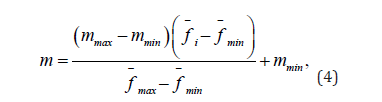
where m – is the required number of extinct, mmax – is the maximum number of extinct, mmin – is the minimum number of extinct. Thus, individuals in less adapted countries will die out more actively than in more adapted ones, which again seems logical, as with reproduction. The algorithm also contains four artificial actions on countries – movement to the leader, exchange of individuals, war and epidemic.
Movement Towards the Leader
Movement towards the leader is an analogue of certain internal political movements that take place in a country, when residents follow the ideals of their leader. This mechanism is inspired by the migration and through this process, all individuals within the country move towards what is considered the best according to the formula (5):
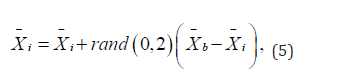
where 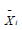 is the current individual,
is the current individual, 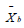 is the best individual in
the country. The mechanism of movement towards the leader will
bring all individuals within the country closer to the best, in order
to find a solution in its vicinity.
is the best individual in
the country. The mechanism of movement towards the leader will
bring all individuals within the country closer to the best, in order
to find a solution in its vicinity.
Exchange of Individuals
The exchange of individuals is an analogue of peaceful relations between different countries, for example, the opening of embassies or migrations between countries, during which countries change k random individuals among themselves. If the size of one of the countries is less than the specified k, then it is equal to half of the number of individuals in that country. This mechanism is largely based on the classical island algorithm and it is assumed that it will reduce the degradation of individual countries and, possibly, prevent the “subsidence” of weak countries by introducing more adapted individuals into them.
War
War is a more radical mechanism than the exchange of individuals, and is an analogue of real military conflicts with the extermination of the enemy and the capture of prisoners. War not only controls the degeneration of individual countries, but also limits the total number of individuals within all countries. During the war, two countries choose l random warriors from their set of individuals. The warriors are compared in pairs with the individuals of the opponent’s country and the losers are removed during this comparison. In this case, the number of victories is calculated when comparing for each country. The country that has won more victories takes both its warriors and the enemy’s warriors, in case of a draw, the warriors are returned to their countries. If the size of one of the countries is less than the specified l, then it is equal to the number of individuals in that country.
Epidemic
Epidemic is an action that is primarily aimed at avoiding the degeneration of a country, as well as reducing the total number of individuals. This mechanism assumes that the more epidemics an individual has experienced, the less it is affected by the next epidemic, due to the developed immunity. During the epidemic d% of individuals dies out, e% of individuals is left unaffected, and the rest mutate according to the formula (6):
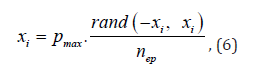
where nep – is the number of epidemics that the current individual has experienced.
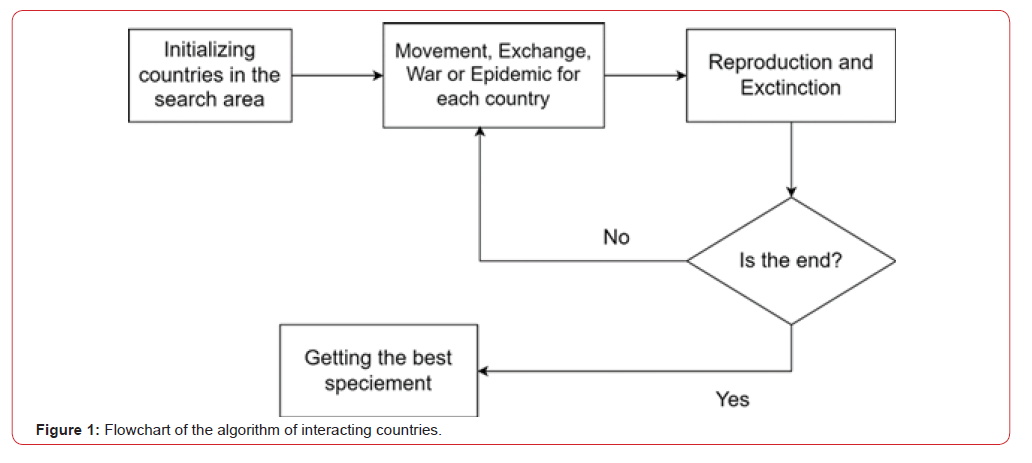
The interacting country algorithm works as follows (Figure 1):
1) The algorithm parameters that are necessary for the
implementation of all its parts are being set.
2) M countries with N individuals are initialized, while each
country is initialized in a separate subdomain of the global
parameter search area.
3) For each country, one artificial action is chosen –
movement to the leader, exchange, war or epidemic. At the
same time, for actions, where a country needs a second country
(exchange, war), the same action is also selected for the second
country.
4) The selected action is performed for each country.
5) Reproduction happens. If, at the time of reproduction,
there is only one individual in the country, this individual joins
a random country, and the previous country of this individual
ceases to exist.
6) Extinction happens.
7) If there are countries without individuals, they are
removed.
8) Steps 3-7 are taken iteratively until the iteration limit is
reached.
9) The best solution is chosen in the best country (Figure 1).
The algorithm of interacting countries was implemented as a part of a library for building the pharmacokinetic models Py Pharm [21], which is written in Python and was previously applied in fitting the parameters of the pharmacokinetic model of intravenous bolus administration of [3H]-cycloprolylglycine [19, 22].
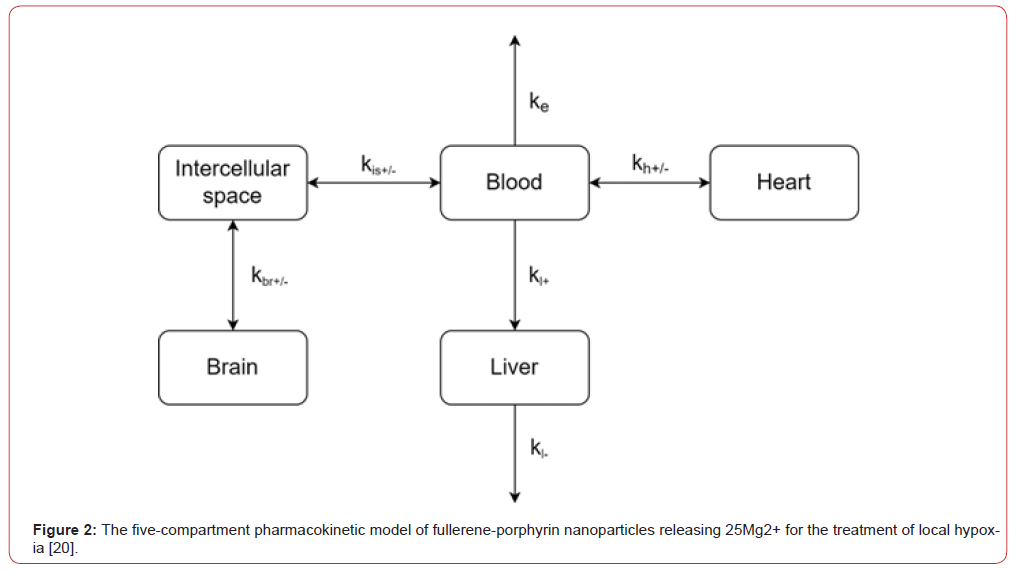
Figure 2:The five-compartment pharmacokinetic model of fullerene-porphyrin nanoparticles releasing 25Mg2+ for the treatment of local hypoxia [20].
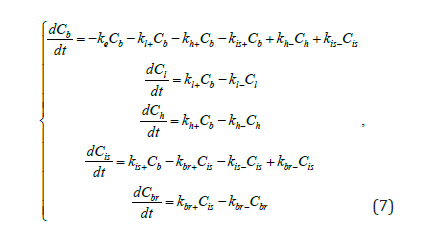
where Cb, Cl, Ch, Cis, Cbr reflect the concentrations of nanoparticles in blood, liver, heart, brain intercellular space and brain cells, respectively.
Function (8) was used to quantify the accuracy of the model, as well as an objective function for fitting model parameters using the interacting countries algorithm:
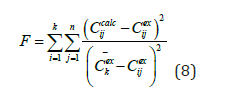
where k is the number of data sets, n is the number of data
in the i-th set, 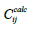 is a calculated concentration value for the i-th
set at j-th point and
is a calculated concentration value for the i-th
set at j-th point and 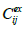 is the experimental one,
is the experimental one, 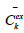 is a mean
concentration value over the i-th experimental data set.
is a mean
concentration value over the i-th experimental data set.
The data sets used were the mean values of PMC16 concentrations in organs - heart, liver and brain cells of the animal, and blood (total 4 data sets) at time points t = 0.25, 0.5, 1, 4, 8, 24 h, when administered once at a dose of 0.2 mg/μg. In detail, the data of the in vivo experiment are given in [20] (number of animals n = 5).
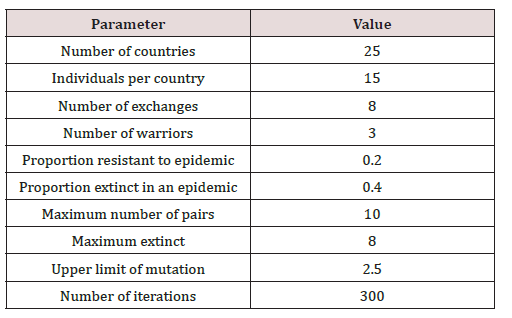
The settings of algorithm of interacting countries (Table 1) were chosen based on the previous studies. These include testing the efficiency of algorithm on a set of test optimisation functions and using the algorithm to find model parameters for the pharmacokinetic model of intravenous bolus administration of [3H]-cycloprolylglycine [19,22].
Results
Fitting The Parameters of the Pharmacokinetic Model of [25 Mg2+] Porphyrin-Fullerene Nanoparticles
In [20], the authors conducted in silico modeling of the pharmacokinetics of [25 Mg2+] porphyrin-fullerene nanoparticles in rats after a single intravenous injection. The authors proposed a five-compartment (Figure 2) model to describe the distribution of a drug substance in the body targeted for the treatment of ischemic stroke. Nanoparticles after administration spread through the blood and enter the organs. Model parameters with a “+” sign reflect the process of substance delivery to an organ, and parameters with a “-” sign reflect the removal of the substance from the organ. For example, kh+ characterizes the rate of PMC16 delivery to the heart, and kh- is responsible for the removal of the substance from the heart back into the blood, Parameter ke is responsible for the elimination of the substance from the blood, and kl- is responsible for its excretion through the liver. Since PMC16 penetrates at a different rate into the stroke region than into the healthy brain region, the introduction of the “intercellular space” compartment allows us to take into account the magnitude of brain damage in ischemic stroke through the kis+ parameter [20].
This model can be described with a system of ordinary differential equations (ODEs) (7) [20].
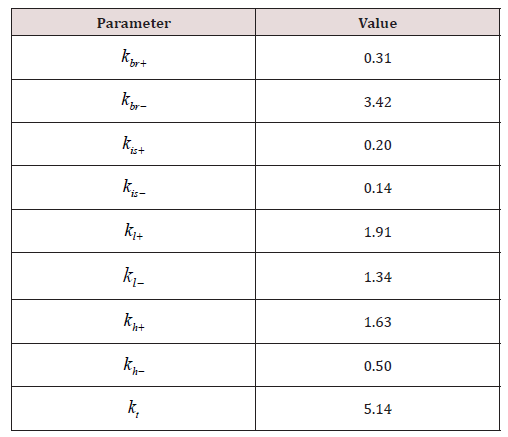
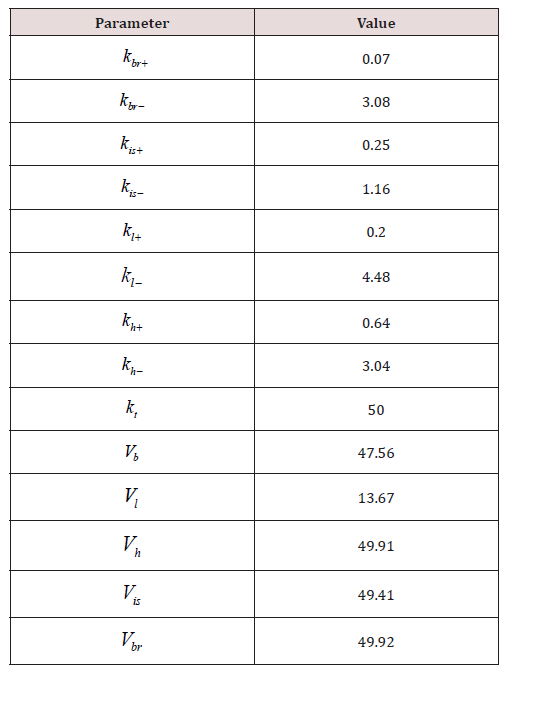
Using this function, the accuracy of the model with parameters presented in the authors’ article [20] was evaluated. The value of the function F was 4.02. In the next step, parameter fitting for the model (7) was conducted with a slight modification. Similar to [20], the elimination constant ke was fixed at the value of 0.077, however, an additional parameter kt was introduced, responsible for converting the input concentration units to output concentration units. This parameter is necessary because the administered dose of the drug is given in mg/ml, the calculations are performed in the same units, while the experimental data on concentrations in organs [20] are provided in ng/mg of protein S125. The fitted values are given in (Table 2).
The model with parameters fitted using the algorithm of interacting countries significantly outperformed the accuracy of the model with original parameter values (F=1.35 vs F=4.02).
At the second step, an attempt to improve the results was made by taking into account the compartment volumes. The second model can also be represented using a system of ODEs (9):
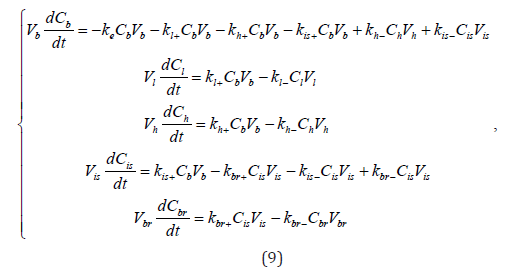
Five new unknowns representing compartment volumes, were added in the second model. Parameters of the second model accounting for compartment volumes was also fitted using our algorithm (Table 3).
Discussion
The second model allowed to improve the value of the objective function, but this improvement was insignificant (F=1.31). The comparison of models’ results is presented on (Figures 3-5), which shows the evolution of drug concentration for specific organs in time. For the original experimental data, confidence intervals were constructed with the confidence level β=0.95. Since the original experimental studies were conducted using n = 5 animals [20], t-distribution for the corresponding mean values was used to construct the confidence intervals. The confidence intervals determination was made using the stats module of SciPy Python library.
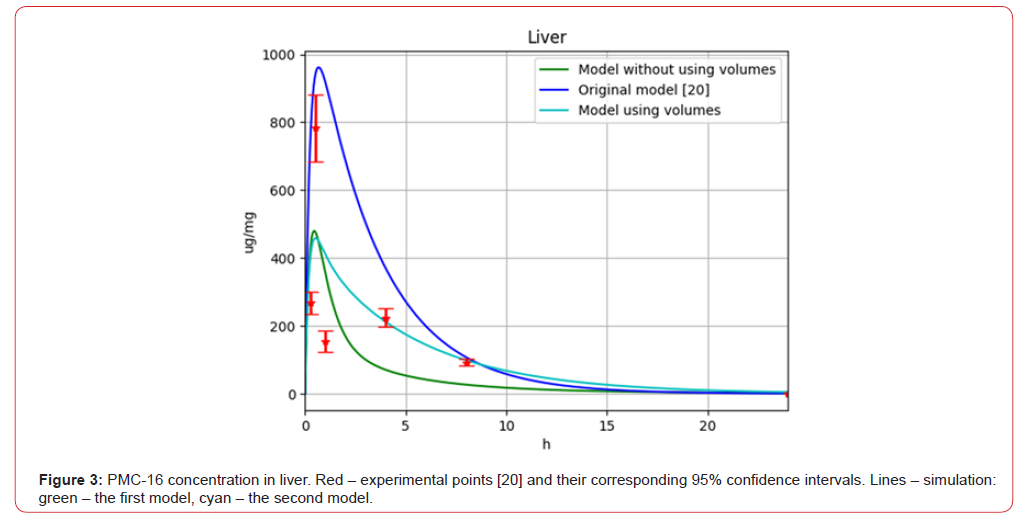
Figure 3:PMC-16 concentration in liver. Red – experimental points [20] and their corresponding 95% confidence intervals. Lines – simulation: green – the first model, cyan – the second model.
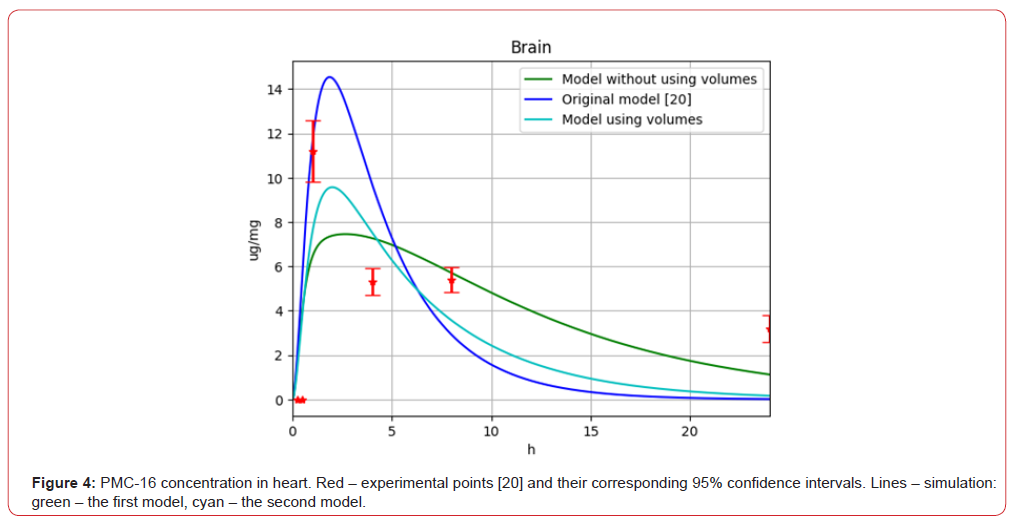
Figure 4:PMC-16 concentration in heart. Red – experimental points [20] and their corresponding 95% confidence intervals. Lines – simulation: green – the first model, cyan – the second model.
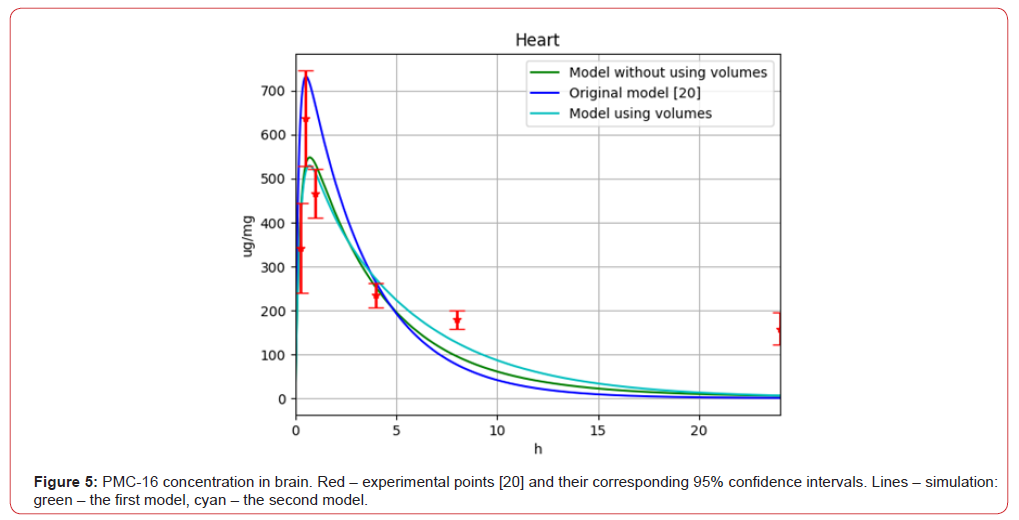
Figure 5:PMC-16 concentration in brain. Red – experimental points [20] and their corresponding 95% confidence intervals. Lines – simulation: green – the first model, cyan – the second model.
Analysing the modelling results, it can be concluded that both our models with parameters fitted the algorithm of interacting countries significantly outperform the original model from [20]. The value of the objective function decreased by approximately 67% for the second model. Meanwhile, it was found that a second model considering compartment volumes slightly better fits the experimental data than a model without compartment volumes. It is also worth noting that none of the three models fully fit within the 95%-confidence intervals for the data, which may indicate either specificity of data or imperfect choice of multi-compartment model structure.
Conclusions
In this article, a new evolutionary algorithm was employed for the first time to tackle pharmacokinetic problems of a fullereneporphyrin nanopharmacophore, PMC16. The algorithm enabled finding pharmacokinetic parameters of two variants of a fivecompartment model for [25Mg2+] porphyrin-fullerene nanoparticles (PMC16) after their single intravenous administration. Results obtained by the algorithm surpassed the accuracy of our previous work [20] approximately threefold, besides proving its efficiency for models with varying numbers of unknowns. Consequently, an evolutionary algorithm can effectively be applied in the pharmacokinetic modelling for promising nanopharmacophores, and may be further used in solving various pharmacokinetic problems.
Authors’ Contributions
Roman S. Krasheninnikov - algorithm and model development, software development, calculations, text, editing; Ivan I. Mitrichev - text, editing, model development, general supervision, Daria I. Zinchenko – data processing, interpretation of results, Valentin V. Fursov – text, editing, general supervision, interpretation of results, biomedical aspects, biochemical aspects.
Authors’ Contributions
Roman S. Krasheninnikov - algorithm and model development, software development, calculations, text, editing; Ivan I. Mitrichev - text, editing, model development, general supervision, Daria I. Zinchenko – data processing, interpretation of results, Valentin V. Fursov – text, editing, general supervision, interpretation of results, biomedical aspects, biochemical aspects.
Ethics Approval and Consent to Participate
Studies on humans and animals have not been conducted. Compliance with ethical standards is not required.
Human and Animal Rights
No animals/humans were used in this research.
Consent For Publication
Not applicable.
Availability of Data and Materials: Any data is available on e-mail also upon request. PyPharm software is available at https://pypi.org/project/pypharm/
Funding
Funding
Conflicts of Interest
The authors declare no conflict of interest financial or otherwise.
Acknowledgements
Prof. Dr. Dmitry A. Kuznetsov, Prof. Dr. Eleonora M. Koltsova, Moscow, Russian Federation, are to be sincerely thanked for their kind assistance with preparation of a manuscript, consultations and for their encouraging comments on this work.
References
- De Jong KA (2006) Evolutionary computation: a unified approach. MIT Press.
- Eiben AE, Smith JE (2006) Introduction to Evolutionary Computing. Natural Computing Series.
- AL Salami NMA (2009) American Journal of Engineering and Applied Sciences. 2: 789-795.
- Vikhar PA (2016) International Conference on Global Trends in Signal Processing, Information Computing and Communication (ICGTSPICC). IEEE: 261-265.
- Sanchez E, Squillero G, Tonda A (2012) Industrial applications of evolutionary algorithms. Intelligent systems reference library 34: i-120.
- Habib M, Mirjalili S, Faris H, Aljarah I (2020) Evolutionary machine learning techniques: algorithms and applications (Algorithms for intelligent systems) Springer: 175-201.
- Goranova M, Ochoa G, Maier P, Hoyle A (2022) Evolutionary optimisation of antibiotic dosing regimens for bacteria with different levels of resistance. Artificial Intelligence in Medicine 133: 102405.
- Yamashita F, Fujita A, Sasa Y, Higuchi Y, Tsuda M, et al. (2017) An Evolutionary Search Algorithm for Covariate Models in Population Pharmacokinetic Analysis. Journal of Pharmaceutical Sciences 106: 2407-2411.
- Morozov M, Nuske E, Serra HA (2021) The use of GA-RxODE (Genetics Algorithms and Running simulations from Ordinary Differential Equations-based model) method to optimize bioequivalence studies. J Pharm Innov 16: 152-159.
- Sherer EA, Sale ME, Pollock BG, Belani CP, Egorin MJ, et al. (2012) Application of a single-objective, hybrid genetic algorithm approach to pharmacokinetic model building. J Pharmacokinet Pharmacodyn 39(4): 393-414.
- Sepúlveda C, Melin P, Castillo O, Kacprzyk J, Reformat M, et al. (2018) In Fuzzy Logic in Intelligent System Design: Theory and Applications; Advances in Intelligent Systems and Computing 648: 214-221.
- Saltzman WM (2001) Drug delivery: engineering principles for drug therapy. Topics in chemical engineering; Oxford University Press.
- Amirshahi N, Alyautdin RN, Sarkar S, Rezayat SM, Orlova MA, et al. (2008) Porphyrin-fullerene nanoparticles for treatment of hypoxic cardiopathies. Nanotechnol Russia 3: 611-621.
- Sarkar S, Rezayat S, Buchachenko A, Kuznetsov D, Orlova M, et al. (2008) New water soluble porphylleren compounds.
- Sarkar S, Rezayat S, Buchachenko A, Kuznetsov D, Orlova M, et al. (2008) Use of a magnesium isotope for treating hypoxia and a medicament comprising the same.
- Buchachenko AL, Kouznetsov DA, Arkhangelsky SE, Orlova MA, Markarian AA (2005) CBB. 43: 243-252.
- Buchachenko AL, Kouznetsov DA, Arkhangelsky SE, Orlova MA, Markarian AA (2005) Spin biochemistry: intramitochondrial nucleotide phosphorylation is a magnesium nuclear spin controlled process. Mitochondrion 5(1): 67-69.
- Buchachenko AL, Kouznetsov DA, Orlova MA, Markarian AA (2005) Proc. Natl. Acad. Sci. U.S.A. 102: 10793-10796.
- Krasheninnikov RS, Mitrichev II (2023) Applied Mathematics and Control Sciences. 4: 126-135.
- Fursov VV, Zinchenko DI, Namestnikova DD, Kuznetsov DA (2022) Bulletin of RSMU. 4: 58-63.
- (2024) pypharm: Module for solving pharmacokinetic problems. URL: https://pypi.org/project/pypharm/ (Accessed 27.03.2024).
- Kovalev GI, Zolotarev Yu A, Dadayan AK, Shram SI, Abdullina АА, et al. (2018) Pharmacokinetics and pharmacodynamics 3: 48-59.



 We use cookies to ensure you get the best experience on our website.
We use cookies to ensure you get the best experience on our website.