Research Article 
 Creative Commons, CC-BY
Creative Commons, CC-BY
Gene Associated Divergence of COVID-19 Morbidity & COVID-19 Vaccines
*Corresponding author: Jomana Al-Azzam, Department of Biotechnology & Genetic Engineering, Faculty of Science, Philadelphia University, Jordan.
Received: July 19, 2021; Published: September 01, 2021
DOI: 10.34297/AJBSR.2021.14.001955
Abstract
The Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2), also known as COVID-19, drastically changed our everyday lives. It has indeed caused a considerable increase in morbidity and mortality rates worldwide.
There is an underlying complex interplay between the infectious agents and the human host, which is related to different biological mechanisms. Phenotypic spectrums associated with SARS-CoV-2 infection or COVID-19 range from asymptomatic to severe systemic complications such as pneumonia, respiratory failure, and death. Around 15 % of cases are severe. Some are accompanied by a dysregulated immune system or a cytokine storm, and others with both.
There is increasing evidence that the severe manifestations of COVID-19 might attribute to human genetic variants. Polymorphisms in genes that are related to immune deficiency and or inflammasome activation (cytokine storm) are examples of these variants.
The question is, is the variability of the hosts’ genetic background the reason behind the different responses to COVID-19? Or are there other factors? Case reports and GWAS studies showed that the susceptibility to severe viral infections was associated with the genetic variants in the immune response genes. Identifying the candidate’s genes is likely to aid in explaining why COVID-19 symptoms are severe to some but not others. Not to mention that it will provide insights that help us further understand the pathogenesis of severe COVID-19, to then make it possible to come out with more effective treatments and vaccines.
There are forty genes associated with viral susceptibility. Twenty-one of them were associated with severe SARS-CoV disease and severe COVID-19. Some of these genes are implicated in the TLR pathways and others in C-lectin pathways. Additionally, there are some genes related to inflammasome activation (cytokine storm). This explains the diverse observations of the disease’s severity associated with genes.
A vaccine should protect humans against SARS-CoV-2 severe infection and death. It acts against infection, disease, or transmission. A vaccine capable of reducing any of these elements will contribute to disease control. Vaccines should be safe, effective, immunogenic, economic, and upscalable.
Brief Historical Perspective
Part of inter-individual differences is attributable to the host genetics profile. [1][2] Mendelian disorders causing deficiencies of the immune response in the human host to viruses (reviewed in Casanova JL et al. 2020 [3] gave a base to search for the influence of germline genetic variations on sporadic infectious diseases.
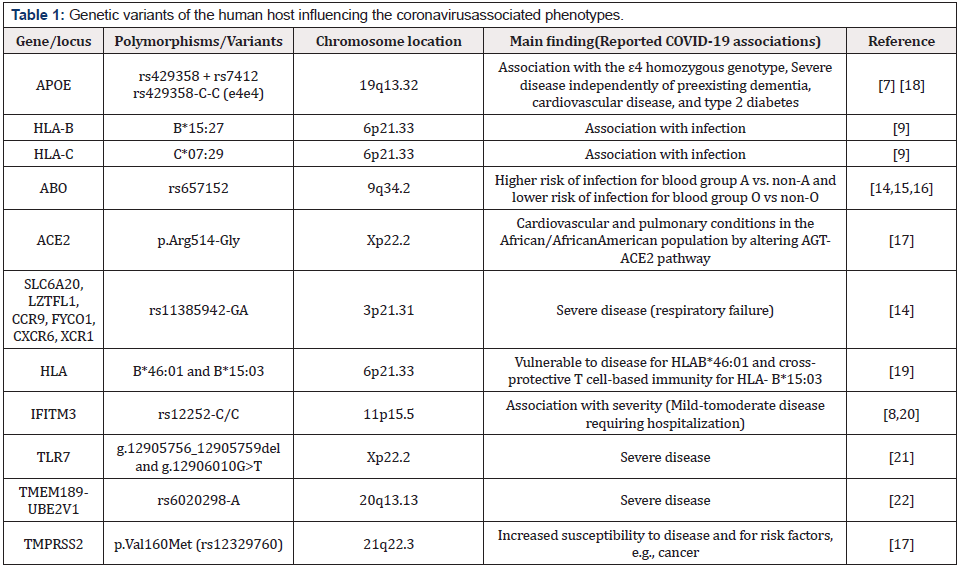
The current paradigm for complex disorders implies that the genetic susceptibility to infection is not Mendelian-but may be either monogenic or polygenic-and postulates that any single genetic biomarker may be associated with a risk with low to very slight effect. SARS-CoV-2 receptor-binding spike motif binds to the human Angiotensin-Converting Enzyme 2 (ACE2), triggering a cascade of events leading to the fusion between the cell and viral membranes for cell entry. So, a hypothesis arose that the variability in ACE2 (and other factors) structure and expression is related to different susceptibility to COVID-19 (recently reviewed by Devaux CA et al. [4], connections between disease progression and age, sex, immune-sufficiency, and having comorbidities, are established with vulnerable individuals (Figure 1).
Primary viral replication occurs when the virus reaches the mucosal epithelium of the upper respiratory tract, SARS-CoV-2 can reach lung epithelial cells where further viral replication occurs.
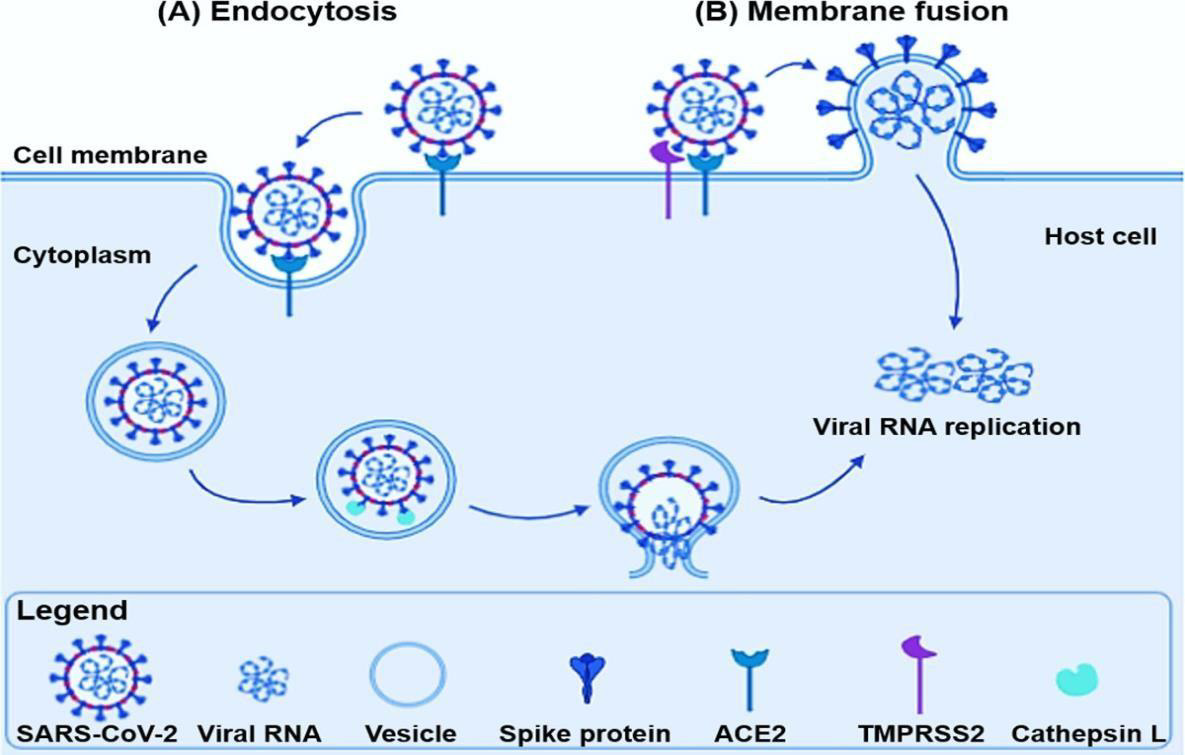
SARS-CoV-2 binds via its Spike (S) protein to the ACE2 receptor in the lungs (same mechanism as in airway epithelial cells) and TMPRSS2 (increases viral entry to the host). The binding interaction promotes cellular endosomal entrance via cathepsin-Ldependent endocytosis. Host proteases cleave S protein, which activates it to trigger the process of membrane fusion [5,6].
The virus can enter cells either by direct cellular entry (membrane fusion then injection of the viral genome into the host’s cytoplasm) or endocytosis (material enters the host after being surrounded by an area of the cell membrane then buds off inside the cell to form a vesicle). Once inside the cell, viral-specific RNA and proteins are synthesized within the cytoplasm [7-10]. More viral proteins are then assembled from the information within the viral RNA using the host’s cellular machinery, specifically the Golgi apparatus and endoplasmic reticulum [11]. New variants are then assembled by fusing to the plasma membrane and being released as vesicles by the cellular exocytic secretory processes. [12]
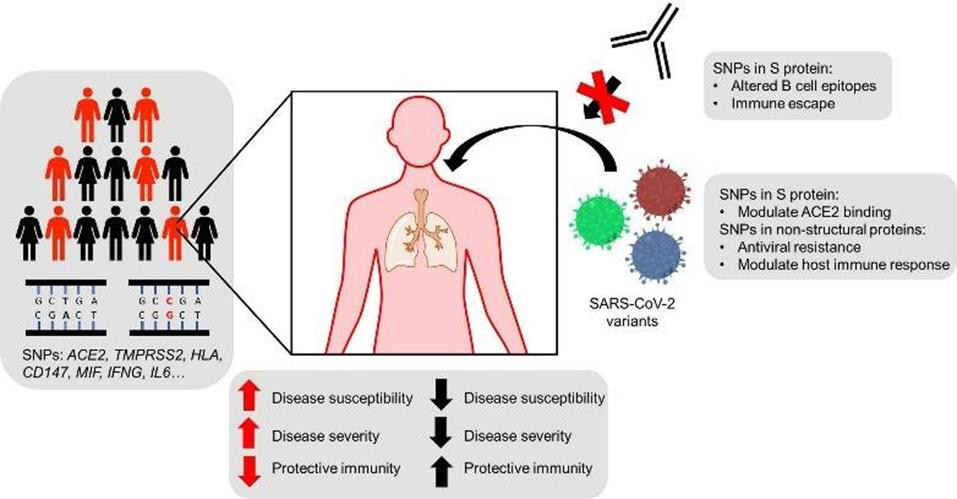
Host genetics and viral variation have a huge impact on SARS‐CoV‐2 infection and COVID‐19 severity. Single nucleotide polymorphisms (SNPs) are present in individuals across a variety of genes` such as ACE2, TMPRSS2, HLA, CD147, MIF, IFNG, IL6) which can be involved in SARS‐CoV‐2 pathology and immunology, as well as other genetic variants that may be able to: Modulate disease susceptibility, increase or decrease disease severity, alter the variety of symptoms developed, and Affect the magnitude and or quality of the immune responses against SARS‐CoV‐2.
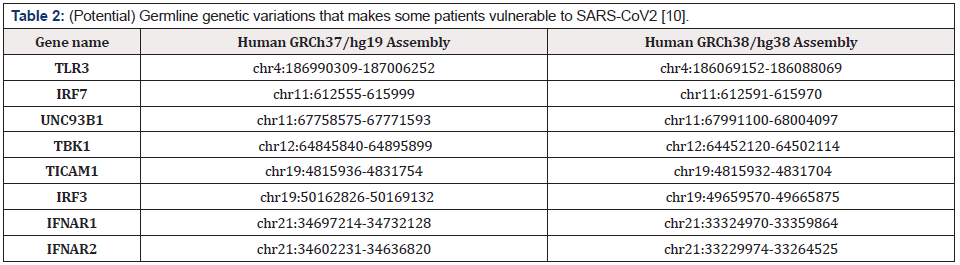
Genetic variants of SARS‐CoV‐2 in addition to host genetic variation show differences in biological activity. examples include the capability of SARS-CoV2 to suppress the activation of the innate immune system by Single amino acid mutations in the spike glycoprotein (which can modulate ACE2 binding or alter B cell epitopes to promote immune escape or render monoclonal antibodies ineffective), it is also able to promote the development of resistance to antivirals, alter T cell epitopes, disrupt cell-mediated immunity, and modulate host cellular interactions with viral particles in the presence of mutations in non‐structural/accessory proteins [13] (Figure 2) (Table 1 & 2).
Epigenetic Mechanisms Regulating COVID-19 Infection
Global DNA methylation, ACE2 gene methylation and posttranslational histone modifications drive differences in host tissue-, biological age- and sex-biased patterns of viral infection. Epigenetic changes impact genome stabilization, maintenance of cellular homeostasis, and affect the pathophysiology of the viral infection [14].
COVID-19 and Age
By far, age is the strongest predictor of the risk of dying for an infected person, measured by a metric, infection fatality ratio (IFR), in which the proportion of people infected with the virus, including those who didn’t get tested or show symptoms, to the people who will die as a result of the infection [15-17]. Age and disease severity are correlated with multiple immunological characteristics [18- 21]. There are correlations between low frequencies of naive CD8 +, CD4+ T cells, age and COVID-19 disease severity. Coordinated CD4+ T cell, CD8+ T cell, and antibody responses are (protective), but uncoordinated responses fail to control the disease [22-25]. This discovery Formed a connection between aging, impaired adaptive immune responses, and dysregulated immune responses to SARSCoV- 2 [26-28].
COVID-19 and Sex
SARS-CoV-2 binds to the ACE2, present in high levels in the testis. The development of acute respiratory distress syndrome (ARDS) is associated with elevated IL-6. Low T is associated with ARDS [29].
Almost half of the admitted COVID-19 patients developed ARDS, and half of them died. Elderly male patients are more likely to develop ARDS and die [30-32]. Low T levels may worsen the severity of COVID-19 infection in older males. Normal T levels may offer some protection against COVID-19. [24, 33] females are endowed with a higher capacity to produce antibodies, which correlates with viral clearance and disease resolution in the context of SARS-Cov-2 infection.
Many X-linked immune genes escape X inactivation, showing bi-allelic expression in female immune cells, particularly in plasmacytoid dendritic cells (pDCs). PDCs are more active in females and endowed with high capability to induce IFN-α-mediated B cell activation and differentiation into antibody-producing plasma cells throughout epigenetic mechanisms linked to trained immunity. It was hypothesized that following SARS-CoV-2 infection, epigenetic modifications of X-linked genes involved in pDC-mediated type I IFN (IFN-I) signaling occurs more effectively in females, for inducing neutralizing antibody response as an immune correlate driving sex-biased disease outcome [30]. Differences in innate and adaptive immune responses, genetic factors, and an interplay between sex hormones and immune effectors, as well as gender-specific behavior differences, particularly the immunological divergence in response to viral infection, could potentially influence not only COVID-19 pathogenesis and disease course, but also the response to antiviral drugs and vaccines [32]. This might explain why severe COVID-19 and mortality rates are higher in men than women.
COVID-19 and the Immune System
The first line of defense when a pathogenic invader enters the body is the immune system. The main course of COVID-19 disease occurs because of a misbalance of the immune response, often leading to misregulation and worsening of the infection [34]. Examples of candidate genes that have the potential to play a role in the immune defense against COVID-19 (might explain the variable prevalence of the infection): complement component 3 (C3), galactoside 2-Alpha-L-fucosyltransferase 2 (FUT2), haptoglobin (Hp), vitamin D binding protein (DBP), human homeostatic iron regulator protein (HFE), cystic fibrosis transmembrane conductance regulator (CFTR), and angiotensin-converting enzyme 1 (ACE1).
Univariate studies showed that ACE1 D/I, C3, CFTR, and HFE polymorphisms correlated significantly with COVID-19 prevalence/ mortality, while multivariate analysis, only ACE1 D/I and C3 polymorphisms were determinants for COVID-19 prevalence/ mortality. Whereas ACE1 D/I polymorphism shows functional links with ACE2 in COVID-19, C3 acts as a critical step in the virusinduced inflammation [30].
Molecular Grade Vaccines
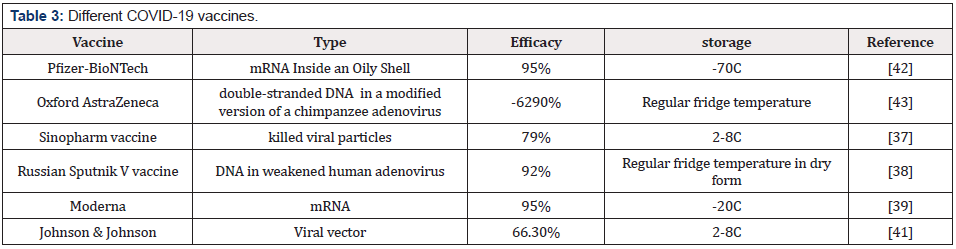
An efficacious vaccine is necessary to prevent the increase in morbidity and mortality. Safety and immunogenicity are the most important categories to look for in a vaccine. The vaccine development goal is to gain direct evidence of vaccine efficacy in protecting humans against SARS-CoV-2 infection. The vaccine should be capable of reducing infection, disease, or transmission and induce the best immunogenic reaction in safe manners to be considered a candidate vaccine that contributes to disease control. Protection against severe disease and death is the most vital efficacy endpoint [35]. Each vaccine exhibits a different potency and duration of efficacy, as determined by the antigen design, adjuvant molecules, vaccine delivery platforms, and immunization method [36]. Most COVID-19 vaccines use a nucleic acid segment (mRNA or DNA) that encodes for the viral S (spike) protein of SARS-CoV2, translated by host cells and the product gets recognized by the immune system (and antibodies are built against it) [36-38]. Other vaccines use inactivated viruses for the same purpose. Duration of protection, safety, and mass vaccination are crucial when evaluating any vaccine [39-41] (Table 3).
Opinion on Blood Clotting Caused by Astrazeneca Vaccine
Clotting dysregulation was observed in severe COVID-19 patients [42-45]. Several mechanisms may contribute to immunothrombosis, for instance, elevated levels of pro-inflammatory cytokines [46- 49]. So, I think aside from some additives that supposedly trigger clotting in the AstraZeneca vaccine, patients that would have had a severe case did get immunothrombosis when taken AstraZeneca vaccine being the most immunogenic one.
Possible Treatment Methods
Clinical outcome of COVID-19 in C3S and ACE1 D allele carriers to study the role of C3 and ACE1 D/I polymorphisms in COVID-19 have potential effects on treatment response [44]. drugs enhancing ACE2 activity may become one of the most promising approaches for treating COVID-19 in the future [33]. IL-18, CCR1, CCR9, and EndoU (coronavirus protein) inhibition as a possible treatment for COVID-19.
Summary of Some Established Principles
Angiotensin-converting enzyme 2 (ACE2 “receptor”) interacts with the S protein of SARS-CoV-2 which provides the entry point for the virus to hook into and infect a wide range of human cells [6]. Humans are not equal concerning the expression levels of the cellular ACE2 [4]. ACE2-expressing organs do not equally participate in COVID-19 pathophysiology, implying that other mechanisms are involved in orchestrating cellular infection [27].
Variable responses to coronaviruses (CoVs) are influenced by the variability of the hosts’ genetic background (Emilio Di Maria et al). [5]
Seniors and those with co-morbidities are prone to COVID-19 owing to immunosenescence and exaggerated inflammatory responses and possible treatment with convalescent plasma [40]. The magnitude of neutralizing antibody responses is positively correlated with disease severity (smaller in asymptomatic individuals and decreases faster than in symptomatic). Pre-existing cross-reactive immunity leads to a better prognosis [40].
Different host immune responses to SARS-CoV-2 infection explain why males and females, young and old persons infected with this virus have markedly distinct disease severity [6].
Potential regulators of ACE2 in the human lung, including genes related to histone modifications (e.g. HAT1, HDAC2, and KDM5B) are found in patients with severe COVID-19 [23]. Patients with comorbidities associated with severe COVID-19 found that ACE2 was highly expressed in these patients compared to control individuals [26].
Androgen’s TMPRSS2-mediated actions might explain both the low fatalities observed in prepubertal children and the differences between sexes regarding SARS-COV2 infection. Androgen sensitivity may be a critical factor in determining COVID-19 disease severity, and sensitivity tests can, therefore, help in predicting patient outcomes [29].
References
- Albright Fs, Orlando P, Pavia At, Jackson Gg, Albright Lac (2008) Evidence for a heritable predisposition to death due to influenza. J Infect Dis 197(1): 18-24.
- Sorensen Tia, Nielsen Gg, Andersen Pk, Teasdale Tw (1988) Genetic and environmental influences on premature death in adult adoptees. N Engl J Med 318(12): 727-32.
- Casanova Jl, Su Hc, Effort Chg (2020) A global effort to define the human genetics of protective immunity to SARS-CoV-2 infection. Cell 181(6): 1194-1199.
- Devaux Ca, Rolain Jm, Raoult D (2020) ACE2 receptor polymorphism: Susceptibility to SARS-CoV-2, hypertension, multi-organ failure, and COVID-19 disease outcome. J Microbiol Immunol Infect 53(3): 425-435.
- Emilio Di Maria, Andrea Latini, Paola Borgiani, Giuseppe Novelli (2020) Genetic variants of the human host influencing the coronavirus-associated phenotypes (SARS, MERS and COVID-19): rapid systematic review and field synopsis.Hum Genomics 14(1): 30.
- Meng-Yuan Li, Lin Li, Yue Zhang, Xiao-Sheng Wang (2020) Expression of the SARSCoV-2 cell receptor gene ACE2 in a wide variety of human tissues. Infect Dis Povert 9(1): 45.
- Kuo CL, Pilling LC, Atkins JL, Masoli JAH, Delgado J et al. (2020) genotype predicts severe COVID-19 in the UK Biobank community cohort. The Journals of Gerontology 75(11): 2231-2232.
- Zhang Y, Qin L, Zhao Y, Zhang P, Xu B, Li K et al. (2013) Interferon-Induced Transmembrane Protein 3 Genetic Variant rs12252-C Associated with Disease Severity in Coronavirus Disease 2019. Nat Commun 4: 1418.
- Wang W, Zhang W, Zhang J, He J, Zhu F (2020) Distribution of HLA allele frequencies in 82 Chinese individuals with coronavirus disease-2019 (COVID-19). HlA.
- Science, COVID Human Genetic Effort (Sep 24 2020), (in reference to 659 patients with life-threatening COVID-19 pneumonia relative to 534 asymptomatic patients of various ancestries/ Variants underlying autosomal-recessive/dominant deficiencies were identified in 23 patients (3.5%) 17-77 years of age).
- (2020) New coronavirus variant: what is the spike protein and why are mutations on it important? New coronavirus variant: what is the spike protein and why are mutations on it important?
- (2020) Elsevier Understanding the complexities of SARS-CoV2 infection and its immunology: A road to immune-based therapeutics.
- Inna G Ovsyannikova , Iana H Haralambieva, Stephen N Crooke, Gregory A Poland, Richard B Kennedy (2020) The role of host genetics in the immune response to SARS-CoV-2 and COVID-19 susceptibility and severity. Immunol Rev 296(1): 205-219.
- Ellinghaus D, Degenhardt F, Bujanda L, et al. (2020) Genomewide association study of severe COVID-19 with respiratory failure. NEJM
- Zhao J, Yang Y, Huang H, Et al. (2020) Relationship between the ABO blood group and the COVID-19 susceptibility. Clin Infect Dis.
- Wu Y, Feng Z, Li P, Yu Q (2020) Relationship between ABO blood group distribution and clinical characteristics in patients with COVID-19. Clin Chim Acta 509: 220-223.
- Hou Y, Zhao J, Martin W, Asha Kallianpur, Mina K Chung, et al. (2020) New insights into genetic susceptibility of COVID-19: an ACE2 and TMPRSS2 polymorphism analysis. BMC Med 18(1): 216.
- Kuo Cl, Pilling Lc, Atkins Jl, Jane A H Masoli, João Delgado et al. (2020) APOE e4 genotype predicts severe COVID-19 in the UK Biobank community cohort. J Gerontol A Biol Sci Med Sci 75(11): 2231-2232.
- Nguyen A, David Jk, Maden Sk, Mary A. Wood, Benjamin R. Weeder, et al. (2020) Human leukocyte antigen susceptibility map for severe acute respiratory syndrome coronavirus. J Virol 94(13).
- Thevarajan I, Nguyen Tho, Koutsakos M, Julian Druce, Leon Caly, et al. (2020) Breadth of concomitant immune responses prior to patient recovery: a case report of non-severe COVID-19. Nat Med 26(4): 453-455.
- Van Der Made CI, Simons A, Schuurs-Hoeijmakers J, Guus Van Den Heuvel, Tuomo Mantere, et al. (2020) Presence of genetic variants among young men with severe COVID-19. JAMA 324(7): 663-673.
- Wang F, Huang S, Gao H, Yuwen Zhou, Changxiang Lai et al. (2020) Initial whole genome sequencing and analysis of the host genetic contribution to COVID-19 severity and susceptibility. Cell Discov 6(1): 83.
- Pinto BGG, Oliveira AER, Singh Y, Leandro Jimenez, Andre N A. Gonçalves et al. (2020) ACE2 Expression Is Increased in the Lungs of Patients with Comorbidities Associated with Severe COVID-19.
- Papadopoulos V, Li L, Samplaski M Andrology (2020) why does COVID-19 kill more elderly men than women? Is there a role for testosterone? Andrology 9(1): 65-72.
- (2020) The coronavirus is most deadly if you are older and male - new data reveal the risks, Nature, The coronavirus is most deadly if you are older and male - new data reveal the risks.
- Pinto Bgg, Oliveira Aer, Singh Y, Jimenez L, Gonçalves Ana, et al. (2020) ACE2 Expression Is Increased in the Lungs of Patients with Comorbidities Associated with Severe COVID-19. J Infect Dis 222(4): 556-563.
- Bourgonje Ar, Abdulle Ae, Timens W, Hillebrands Jl, Navis Gj et al. (2020) Angiotensin-converting enzyme 2 (ACE2), SARS-CoV-2 and the pathophysiology of coronavirus disease 2019 (COVID-19). J Pathol 251(3): 228-248.
- Carolyn Rydyznski Moderbacher, Sydney I. Ramirez, Jennifer M Dan, Alba Grifoni, Kathryn M Hastie et al. (2020) Antigen-Specific Adaptive Immunity to SARS-CoV-2 in Acute COVID-19 and Associations with Age and Disease Severity. Cell 183(4): 996-1012.
- Mohamed S Mohamed, Thiago C. Moulin, Helgi B. Schiöth (2020) Sex differences in COVID-19: the role of androgens in disease severity and progression. Endocrine 71(1): 3-8.
- Gabriele L, Fragale A, Romagnoli G, Parlato S, Lapenta C, et al. (2021) Type I IFN-dependent antibody response at the basis of sex dimorphism in the outcome of COVID-19. Cytokine Growth Factor Rev 58: 66-74.
- Joris R Delanghe, Marc L De Buyzere, Marijn M Speeckaert (2021) Genetic Polymorphisms in the Host and COVID-19 Infection Genetic Polymorphisms in the Host and COVID-19 Infection. Adv Exp Med Biol 1318: 109-118.
- Raza HA, Sen P, Bhatti OA, Gupta L Rheumatol Int (2021) Sex hormones, autoimmunity and gender disparity in COVID-19, Springer Sex hormones, autoimmunity and gender disparity in COVID-19.
- Cheng H, Wang Y, Wang Gq (2020) Organ-protective effect of angiotensin- converting enzyme 2 and its effect on the prognosis of COVID- 19. J Med Virol 92(7):726-730.
- Silveira Mm, Moreira Gmsg, Mendonça M (2020) DNA vaccines against COVID-19: Perspectives and challenges. Life Sci 267:118919.
- Hodgson Sh, Mansatta K, Mallett G, Harris V, Emary Krw, et al. (2020) What defines an efficacious COVID-19 vaccine? A review of the challenges assessing the clinical efficacy of vaccines against SARS-CoV-2. Lancet Infect Dis 21(2): e26-e35.
- Park Ks, Sun X, Aikins Me, Moon Jj (2020) Non-viral COVID-19 vaccine delivery systems. Adv Drug Deliv Rev 169:137-151.
- BBC, (2021) Covid: What do we know about China's coronavirus vaccines? Covid: What do we know about China's coronavirus vaccines? - BBC News
- Sputnik V COVID-19 vaccine Sputnik V COVID-19 vaccine - Wikipedia
- Moderna COVID-19 Vaccine Overview and Safety, CBC Moderna COVID-19 Vaccine Overview and Safety | CDC
- Mangalakumari Jeyanathan, Sam Afkhami, Fiona Smaill, Matthew S Miller, Brian D Lichty,et al. (2020) Immunological considerations for COVID-19 vaccine strategies. Nat Rev Immunol 20(10): 615-632.
- Johnson & Johnson’s Janssen COVID-19 Vaccine Overview and Safety Johnson & Johnson’s Janssen COVID-19 Vaccine Overview and Safety | CDC
- How the Pfizer-BioNTech Vaccine Works (2021) How the Pfizer-BioNTech Covid-19 Vaccine Works - The New York Times (nytimes.com)
- How the Oxford-AstraZeneca Vaccine Works (2021)How the Oxford-AstraZeneca Covid-19 Vaccine Works - The New York Times (nytimes.com)
- Genetic Polymorphisms in the Host and COVID-19 Infection. Genetic Polymorphisms in the Host and COVID-19 Infection | SpringerLink
- (2020) Epigenetic mechanisms regulating COVID-19 infection Full article: Epigenetic mechanisms regulating COVID-19 infection (tandfonline.com)
- Middleton Ea, He Xy, Denorme F, Campbell Ra, David Ng, et al. (2020) Neutrophil extracellular traps contribute to immunothrombosis in COVID-19 acute respiratory distress syndrome. Blood 136(10): 1169-1179.
- Morris G, Bortolasci Cc, Puri Bk, Lisa Olive, Wolfgang Marx, et al. (2021) Preventing the development of severe COVID-19 by modifying immunothrombosis. Life Sci 264: 118617.
- Loo J, Spittle Da, Newnham M (2021) COVID-19, immunothrombosis and venous thromboembolism: biological mechanisms. Thorax 76(4): 412-420.
- Nicolai L, Leunig A, Brambs S, Rainer Kaiser, Tobias Weinberger, et al. (2020) Atypon, Immunothrombotic Dysregulation in COVID-19 Pneumonia Is Associated With Respiratory Failure and Coagulopathy. Circulation 142(12):1176-1189.



 We use cookies to ensure you get the best experience on our website.
We use cookies to ensure you get the best experience on our website.